DISCNGINE BLOG
Insights on drug discovery trends, events, product updates, use cases, tech workflows, customer stories, and life at Discngine.
Categories
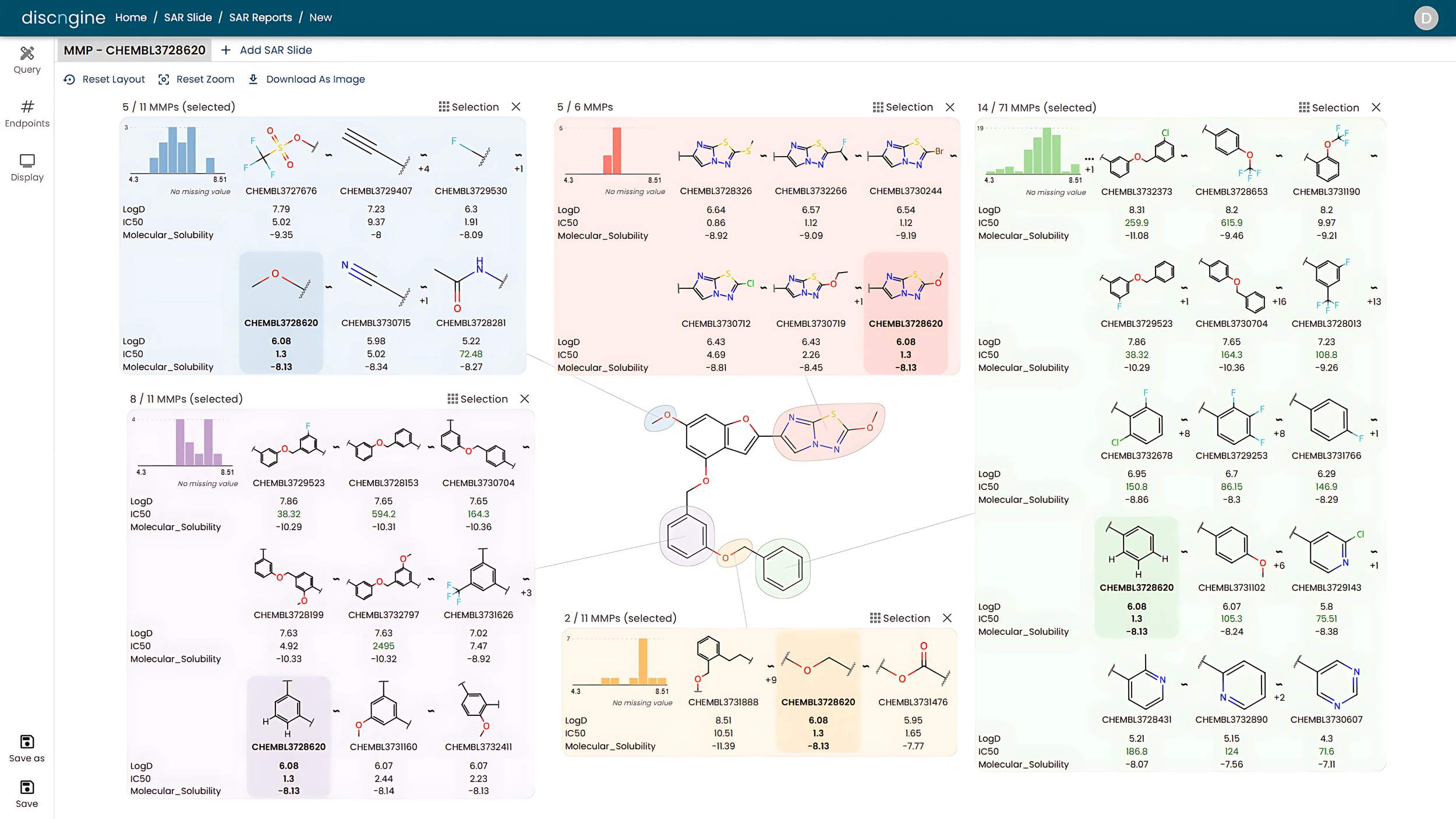
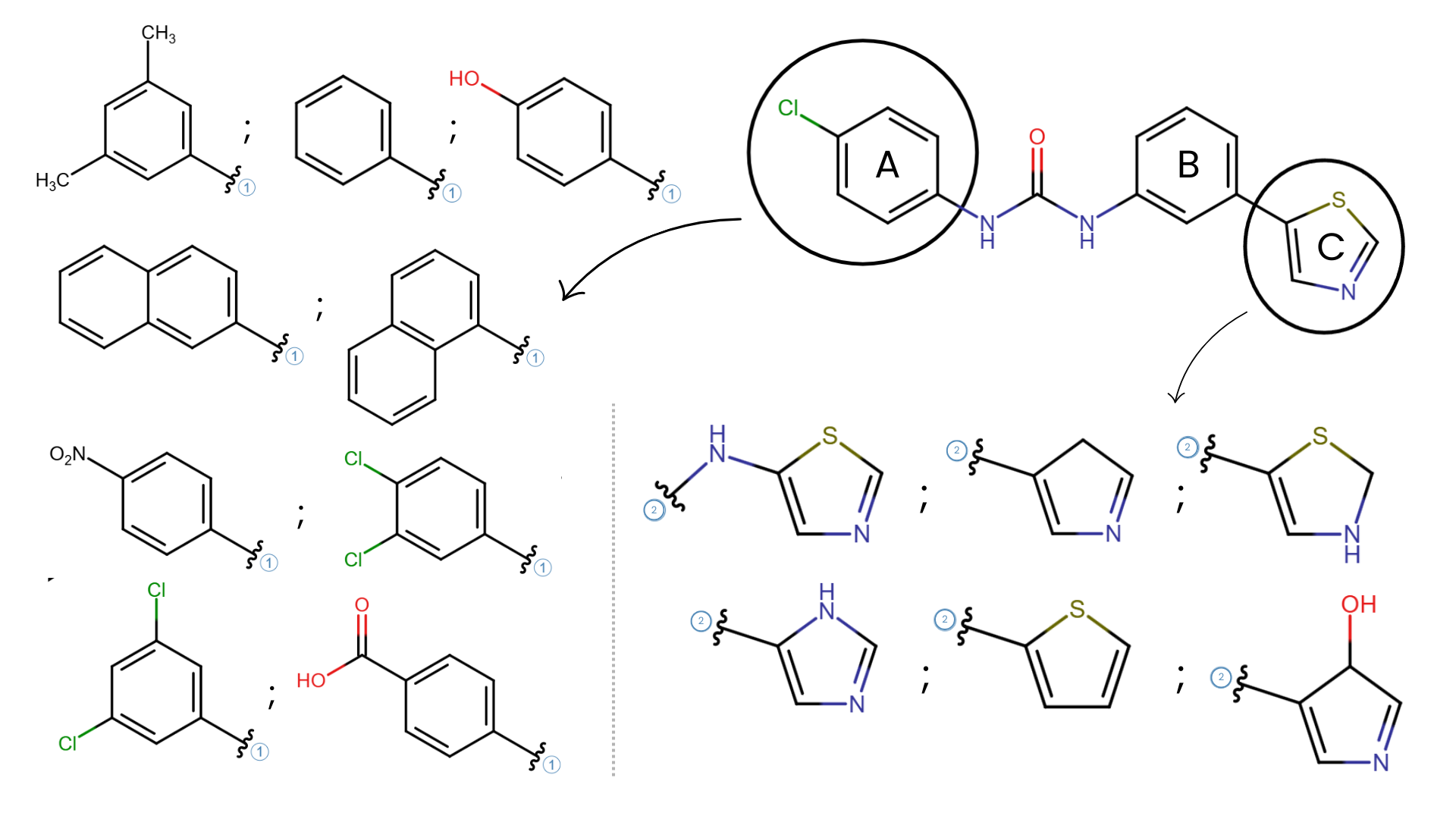
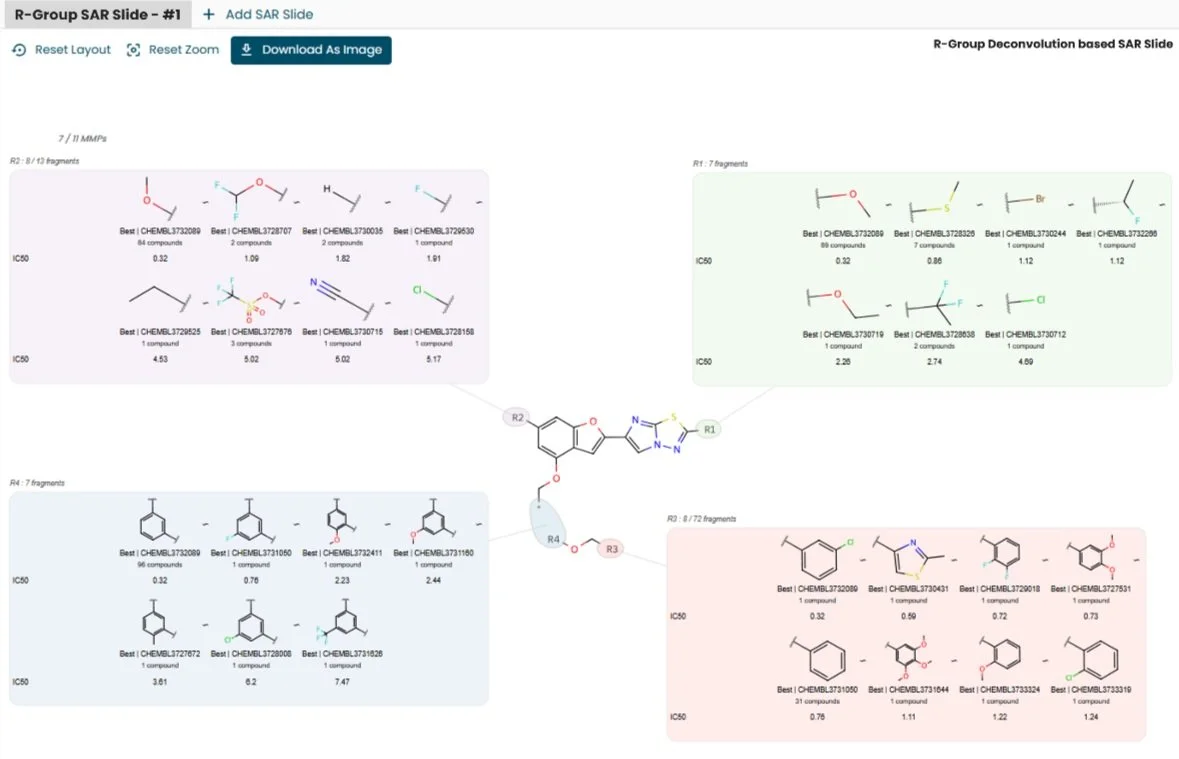
Efficient Structure-Activity Relationship (SAR) Reporting and Exploration in Drug Discovery: Discover Discngine Ideation SAR Slides
Manual structure-activity relationship (SAR) reporting is a time-consuming, tedious, and repetitive task. To regain time from data wrangling, Discngine has developed Ideation SAR Slides - an automated platform for SAR reporting and analysis.
Read more about it here.
Ideation Analytics - SAR Slides Gets Patent Searches with GOSTAR™ Integration
To streamline the Structure-Activity Relationship (SAR) exploration of patents, it is now possible to access GOSTAR™ patent datasets directly within Ideation SAR Slides.
The 3 Hidden Costs of Manual SAR Reporting in Drug Discovery
Structure–activity relationship (SAR) reporting is central to small-molecule drug discovery, yet many teams still rely on manual workflows. This article explores the limitations of manual SAR reporting and how they affect DMTA cycles and decision-making.
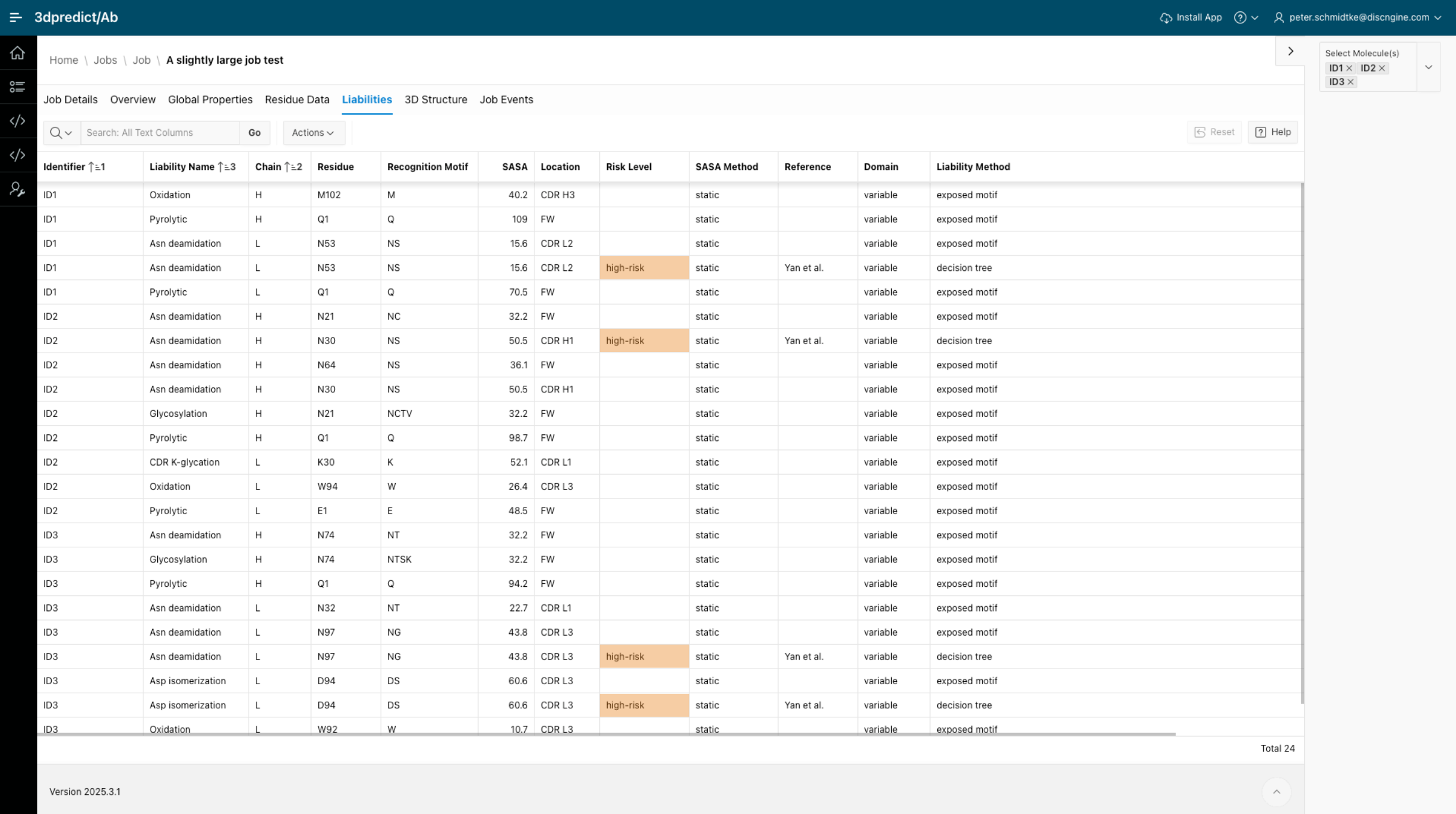
Accelerating early antibody discovery with ensemble-based developability assessment: 3dpredict/Ab
Discover how 3dpredict/Ab tackles current challenges in early antibody discovery by predicting hidden liability risks in candidates through an ensemble-based approach. By looking beyond traditional methods, it provides insights that help researchers make faster, more informed decisions on their candidates.
Highlights from Discngine Meetup Vol. 5: Pushing boundaries in peptide discovery with innovative science and technology
Discngine Meetup celebrated its fifth edition and brought together drug discovery experts to discuss recent progress and challenges in peptide discovery and design. Access event highlights and recordings.
“RNA as a Drug Target” Book Highlight: How targeting RNA emerges as the next frontier for medicinal chemistry
Our scientific expert Riccardo Martini shared an extensive overview of the famous book “RNA as a drug target”, incorporating his thoughts from the modelers' and the medicinal chemists’ perspectives.
Efficient SAR analysis and reporting for rapid compound optimization: A Discngine-Bayer success story
DIscngine collaborated with chemists and data scientists at Bayer Crop Science to overcome the inefficiencies in Structure-Activity Relationship (SAR) report analysis and generation. The part of the new solution is available for commercial use!
Highlights from Discngine Meetup Vol. 4: Biotherapeutic developability assessment strategies
For the fourth year in a row, we have successfully organized our community event with drug discovery experts. Discover the event and access recordings.
Celebrating 20 Years of software for new molecule discovery: Discngine's Milestone Anniversary!
Celebrating 20 years is a significant achievement, to commemorate this significant milestone, we hosted a memorable 20th anniversary party in Paris! The event brought together our valued customers, partners, and other members of our community to celebrate our shared successes and long-lasting collaboration.